Spaces:
Running
on
T4
Running
on
T4
| import os,sys | |
| # install environment goods | |
| #os.system("pip -q install dgl -f https://data.dgl.ai/wheels/cu113/repo.html") | |
| os.system('pip install dgl==1.0.2+cu116 -f https://data.dgl.ai/wheels/cu116/repo.html') | |
| #os.system('pip install gradio') | |
| os.environ["DGLBACKEND"] = "pytorch" | |
| #os.system(f'pip install -r ./PROTEIN_GENERATOR/requirements.txt') | |
| print('Modules installed') | |
| os.system('pip install --force gradio==3.28.3') | |
| os.environ["DGLBACKEND"] = "pytorch" | |
| if not os.path.exists('./SEQDIFF_230205_dssp_hotspots_25mask_EQtasks_mod30.pt'): | |
| print('Downloading model weights 1') | |
| os.system('wget http://files.ipd.uw.edu/pub/sequence_diffusion/checkpoints/SEQDIFF_230205_dssp_hotspots_25mask_EQtasks_mod30.pt') | |
| print('Successfully Downloaded') | |
| if not os.path.exists('./SEQDIFF_221219_equalTASKS_nostrSELFCOND_mod30.pt'): | |
| print('Downloading model weights 2') | |
| os.system('wget http://files.ipd.uw.edu/pub/sequence_diffusion/checkpoints/SEQDIFF_221219_equalTASKS_nostrSELFCOND_mod30.pt') | |
| print('Successfully Downloaded') | |
| import numpy as np | |
| import gradio as gr | |
| import py3Dmol | |
| from io import StringIO | |
| import json | |
| import secrets | |
| import copy | |
| import matplotlib.pyplot as plt | |
| from utils.sampler import HuggingFace_sampler | |
| from utils.parsers_inference import parse_pdb | |
| from model.util import writepdb | |
| from utils.inpainting_util import * | |
| plt.rcParams.update({'font.size': 13}) | |
| with open('./tmp/args.json','r') as f: | |
| args = json.load(f) | |
| # manually set checkpoint to load | |
| args['checkpoint'] = None | |
| args['dump_trb'] = False | |
| args['dump_args'] = True | |
| args['save_best_plddt'] = True | |
| args['T'] = 25 | |
| args['strand_bias'] = 0.0 | |
| args['loop_bias'] = 0.0 | |
| args['helix_bias'] = 0.0 | |
| def protein_diffusion_model(sequence, seq_len, helix_bias, strand_bias, loop_bias, | |
| secondary_structure, aa_bias, aa_bias_potential, | |
| #target_charge, target_ph, charge_potential, | |
| num_steps, noise, hydrophobic_target_score, hydrophobic_potential, | |
| contigs, pssm, seq_mask, str_mask, rewrite_pdb): | |
| dssp_checkpoint = './SEQDIFF_230205_dssp_hotspots_25mask_EQtasks_mod30.pt' | |
| og_checkpoint = './SEQDIFF_221219_equalTASKS_nostrSELFCOND_mod30.pt' | |
| model_args = copy.deepcopy(args) | |
| # make sampler | |
| S = HuggingFace_sampler(args=model_args) | |
| # get random prefix | |
| S.out_prefix = './tmp/'+secrets.token_hex(nbytes=10).upper() | |
| # set args | |
| S.args['checkpoint'] = None | |
| S.args['dump_trb'] = False | |
| S.args['dump_args'] = True | |
| S.args['save_best_plddt'] = True | |
| S.args['T'] = 20 | |
| S.args['strand_bias'] = 0.0 | |
| S.args['loop_bias'] = 0.0 | |
| S.args['helix_bias'] = 0.0 | |
| S.args['potentials'] = None | |
| S.args['potential_scale'] = None | |
| S.args['aa_composition'] = None | |
| # get sequence if entered and make sure all chars are valid | |
| alt_aa_dict = {'B':['D','N'],'J':['I','L'],'U':['C'],'Z':['E','Q'],'O':['K']} | |
| if sequence not in ['',None]: | |
| L = len(sequence) | |
| aa_seq = [] | |
| for aa in sequence.upper(): | |
| if aa in alt_aa_dict.keys(): | |
| aa_seq.append(np.random.choice(alt_aa_dict[aa])) | |
| else: | |
| aa_seq.append(aa) | |
| S.args['sequence'] = aa_seq | |
| elif contigs not in ['',None]: | |
| S.args['contigs'] = [contigs] | |
| else: | |
| S.args['contigs'] = [f'{seq_len}'] | |
| L = int(seq_len) | |
| print('DEBUG: ',rewrite_pdb) | |
| if rewrite_pdb not in ['',None]: | |
| S.args['pdb'] = rewrite_pdb.name | |
| if seq_mask not in ['',None]: | |
| S.args['inpaint_seq'] = [seq_mask] | |
| if str_mask not in ['',None]: | |
| S.args['inpaint_str'] = [str_mask] | |
| if secondary_structure in ['',None]: | |
| secondary_structure = None | |
| else: | |
| secondary_structure = ''.join(['E' if x == 'S' else x for x in secondary_structure]) | |
| if L < len(secondary_structure): | |
| secondary_structure = secondary_structure[:len(sequence)] | |
| elif L == len(secondary_structure): | |
| pass | |
| else: | |
| dseq = L - len(secondary_structure) | |
| secondary_structure += secondary_structure[-1]*dseq | |
| # potentials | |
| potential_list = [] | |
| potential_bias_list = [] | |
| if aa_bias not in ['',None]: | |
| potential_list.append('aa_bias') | |
| S.args['aa_composition'] = aa_bias | |
| if aa_bias_potential in ['',None]: | |
| aa_bias_potential = 3 | |
| potential_bias_list.append(str(aa_bias_potential)) | |
| ''' | |
| if target_charge not in ['',None]: | |
| potential_list.append('charge') | |
| if charge_potential in ['',None]: | |
| charge_potential = 1 | |
| potential_bias_list.append(str(charge_potential)) | |
| S.args['target_charge'] = float(target_charge) | |
| if target_ph in ['',None]: | |
| target_ph = 7.4 | |
| S.args['target_pH'] = float(target_ph) | |
| ''' | |
| if hydrophobic_target_score not in ['',None]: | |
| potential_list.append('hydrophobic') | |
| S.args['hydrophobic_score'] = float(hydrophobic_target_score) | |
| if hydrophobic_potential in ['',None]: | |
| hydrophobic_potential = 3 | |
| potential_bias_list.append(str(hydrophobic_potential)) | |
| if pssm not in ['',None]: | |
| potential_list.append('PSSM') | |
| potential_bias_list.append('5') | |
| S.args['PSSM'] = pssm.name | |
| if len(potential_list) > 0: | |
| S.args['potentials'] = ','.join(potential_list) | |
| S.args['potential_scale'] = ','.join(potential_bias_list) | |
| # normalise secondary_structure bias from range 0-0.3 | |
| S.args['secondary_structure'] = secondary_structure | |
| S.args['helix_bias'] = helix_bias | |
| S.args['strand_bias'] = strand_bias | |
| S.args['loop_bias'] = loop_bias | |
| # set T | |
| if num_steps in ['',None]: | |
| S.args['T'] = 20 | |
| else: | |
| S.args['T'] = int(num_steps) | |
| # noise | |
| if 'normal' in noise: | |
| S.args['sample_distribution'] = noise | |
| S.args['sample_distribution_gmm_means'] = [0] | |
| S.args['sample_distribution_gmm_variances'] = [1] | |
| elif 'gmm2' in noise: | |
| S.args['sample_distribution'] = noise | |
| S.args['sample_distribution_gmm_means'] = [-1,1] | |
| S.args['sample_distribution_gmm_variances'] = [1,1] | |
| elif 'gmm3' in noise: | |
| S.args['sample_distribution'] = noise | |
| S.args['sample_distribution_gmm_means'] = [-1,0,1] | |
| S.args['sample_distribution_gmm_variances'] = [1,1,1] | |
| if secondary_structure not in ['',None] or helix_bias+strand_bias+loop_bias > 0: | |
| S.args['checkpoint'] = dssp_checkpoint | |
| S.args['d_t1d'] = 29 | |
| print('using dssp checkpoint') | |
| else: | |
| S.args['checkpoint'] = og_checkpoint | |
| S.args['d_t1d'] = 24 | |
| print('using og checkpoint') | |
| for k,v in S.args.items(): | |
| print(f"{k} --> {v}") | |
| # init S | |
| S.model_init() | |
| S.diffuser_init() | |
| S.setup() | |
| # sampling loop | |
| plddt_data = [] | |
| for j in range(S.max_t): | |
| output_seq, output_pdb, plddt = S.take_step_get_outputs(j) | |
| plddt_data.append(plddt) | |
| yield output_seq, output_pdb, display_pdb(output_pdb), get_plddt_plot(plddt_data, S.max_t) | |
| output_seq, output_pdb, plddt = S.get_outputs() | |
| yield output_seq, output_pdb, display_pdb(output_pdb), get_plddt_plot(plddt_data, S.max_t) | |
| def get_plddt_plot(plddt_data, max_t): | |
| x = [i+1 for i in range(len(plddt_data))] | |
| fig, ax = plt.subplots(figsize=(15,6)) | |
| ax.plot(x,plddt_data,color='#661dbf', linewidth=3,marker='o') | |
| ax.set_xticks([i+1 for i in range(max_t)]) | |
| ax.set_yticks([(i+1)/10 for i in range(10)]) | |
| ax.set_ylim([0,1]) | |
| ax.set_ylabel('model confidence (plddt)') | |
| ax.set_xlabel('diffusion steps (t)') | |
| return fig | |
| def display_pdb(path_to_pdb): | |
| ''' | |
| #function to display pdb in py3dmol | |
| ''' | |
| pdb = open(path_to_pdb, "r").read() | |
| view = py3Dmol.view(width=500, height=500) | |
| view.addModel(pdb, "pdb") | |
| view.setStyle({'model': -1}, {"cartoon": {'colorscheme':{'prop':'b','gradient':'roygb','min':0,'max':1}}})#'linear', 'min': 0, 'max': 1, 'colors': ["#ff9ef0","#a903fc",]}}}) | |
| view.zoomTo() | |
| output = view._make_html().replace("'", '"') | |
| print(view._make_html()) | |
| x = f"""<!DOCTYPE html><html></center> {output} </center></html>""" # do not use ' in this input | |
| return f"""<iframe height="500px" width="100%" name="result" allow="midi; geolocation; microphone; camera; | |
| display-capture; encrypted-media;" sandbox="allow-modals allow-forms | |
| allow-scripts allow-same-origin allow-popups | |
| allow-top-navigation-by-user-activation allow-downloads" allowfullscreen="" | |
| allowpaymentrequest="" frameborder="0" srcdoc='{x}'></iframe>""" | |
| ''' | |
| return f"""<iframe style="width: 100%; height:700px" name="result" allow="midi; geolocation; microphone; camera; | |
| display-capture; encrypted-media;" sandbox="allow-modals allow-forms | |
| allow-scripts allow-same-origin allow-popups | |
| allow-top-navigation-by-user-activation allow-downloads" allowfullscreen="" | |
| allowpaymentrequest="" frameborder="0" srcdoc='{x}'></iframe>""" | |
| ''' | |
| # MOTIF SCAFFOLDING | |
| def get_motif_preview(pdb_id, contigs): | |
| ''' | |
| #function to display selected motif in py3dmol | |
| ''' | |
| input_pdb = fetch_pdb(pdb_id=pdb_id.lower()) | |
| # rewrite pdb | |
| parse = parse_pdb(input_pdb) | |
| #output_name = './rewrite_'+input_pdb.split('/')[-1] | |
| #writepdb(output_name, torch.tensor(parse_og['xyz']),torch.tensor(parse_og['seq'])) | |
| #parse = parse_pdb(output_name) | |
| output_name = input_pdb | |
| pdb = open(output_name, "r").read() | |
| view = py3Dmol.view(width=500, height=500) | |
| view.addModel(pdb, "pdb") | |
| if contigs in ['',0]: | |
| contigs = ['0'] | |
| else: | |
| contigs = [contigs] | |
| print('DEBUG: ',contigs) | |
| pdb_map = get_mappings(ContigMap(parse,contigs)) | |
| print('DEBUG: ',pdb_map) | |
| print('DEBUG: ',pdb_map['con_ref_idx0']) | |
| roi = [x[1]-1 for x in pdb_map['con_ref_pdb_idx']] | |
| colormap = {0:'#D3D3D3', 1:'#F74CFF'} | |
| colors = {i+1: colormap[1] if i in roi else colormap[0] for i in range(parse['xyz'].shape[0])} | |
| view.setStyle({"cartoon": {"colorscheme": {"prop": "resi", "map": colors}}}) | |
| view.zoomTo() | |
| output = view._make_html().replace("'", '"') | |
| print(view._make_html()) | |
| x = f"""<!DOCTYPE html><html></center> {output} </center></html>""" # do not use ' in this input | |
| return f"""<iframe height="500px" width="100%" name="result" allow="midi; geolocation; microphone; camera; | |
| display-capture; encrypted-media;" sandbox="allow-modals allow-forms | |
| allow-scripts allow-same-origin allow-popups | |
| allow-top-navigation-by-user-activation allow-downloads" allowfullscreen="" | |
| allowpaymentrequest="" frameborder="0" srcdoc='{x}'></iframe>""", output_name | |
| def fetch_pdb(pdb_id=None): | |
| if pdb_id is None or pdb_id == "": | |
| return None | |
| else: | |
| os.system(f"wget -qnc https://files.rcsb.org/view/{pdb_id}.pdb") | |
| return f"{pdb_id}.pdb" | |
| # MSA AND PSSM GUIDANCE | |
| def save_pssm(file_upload): | |
| filename = file_upload.name | |
| orig_name = file_upload.orig_name | |
| if filename.split('.')[-1] in ['fasta', 'a3m']: | |
| return msa_to_pssm(file_upload) | |
| return filename | |
| def msa_to_pssm(msa_file): | |
| # Define the lookup table for converting amino acids to indices | |
| aa_to_index = {'A': 0, 'R': 1, 'N': 2, 'D': 3, 'C': 4, 'Q': 5, 'E': 6, 'G': 7, 'H': 8, 'I': 9, 'L': 10, | |
| 'K': 11, 'M': 12, 'F': 13, 'P': 14, 'S': 15, 'T': 16, 'W': 17, 'Y': 18, 'V': 19, 'X': 20, '-': 21} | |
| # Open the FASTA file and read the sequences | |
| records = list(SeqIO.parse(msa_file.name, "fasta")) | |
| assert len(records) >= 1, "MSA must contain more than one protein sequecne." | |
| first_seq = str(records[0].seq) | |
| aligned_seqs = [first_seq] | |
| # print(aligned_seqs) | |
| # Perform sequence alignment using the Needleman-Wunsch algorithm | |
| aligner = Align.PairwiseAligner() | |
| aligner.open_gap_score = -0.7 | |
| aligner.extend_gap_score = -0.3 | |
| for record in records[1:]: | |
| alignment = aligner.align(first_seq, str(record.seq))[0] | |
| alignment = alignment.format().split("\n") | |
| al1 = alignment[0] | |
| al2 = alignment[2] | |
| al1_fin = "" | |
| al2_fin = "" | |
| percent_gap = al2.count('-')/ len(al2) | |
| if percent_gap > 0.4: | |
| continue | |
| for i in range(len(al1)): | |
| if al1[i] != '-': | |
| al1_fin += al1[i] | |
| al2_fin += al2[i] | |
| aligned_seqs.append(str(al2_fin)) | |
| # Get the length of the aligned sequences | |
| aligned_seq_length = len(first_seq) | |
| # Initialize the position scoring matrix | |
| matrix = np.zeros((22, aligned_seq_length)) | |
| # Iterate through the aligned sequences and count the amino acids at each position | |
| for seq in aligned_seqs: | |
| #print(seq) | |
| for i in range(aligned_seq_length): | |
| if i == len(seq): | |
| break | |
| amino_acid = seq[i] | |
| if amino_acid.upper() not in aa_to_index.keys(): | |
| continue | |
| else: | |
| aa_index = aa_to_index[amino_acid.upper()] | |
| matrix[aa_index, i] += 1 | |
| # Normalize the counts to get the frequency of each amino acid at each position | |
| matrix /= len(aligned_seqs) | |
| print(len(aligned_seqs)) | |
| matrix[20:,]=0 | |
| outdir = ".".join(msa_file.name.split('.')[:-1]) + ".csv" | |
| np.savetxt(outdir, matrix[:21,:].T, delimiter=",") | |
| return outdir | |
| def get_pssm(fasta_msa, input_pssm): | |
| if input_pssm not in ['',None]: | |
| outdir = input_pssm.name | |
| else: | |
| outdir = save_pssm(fasta_msa) | |
| pssm = np.loadtxt(outdir, delimiter=",", dtype=float) | |
| fig, ax = plt.subplots(figsize=(15,6)) | |
| plt.imshow(torch.permute(torch.tensor(pssm),(1,0))) | |
| return fig, outdir | |
| #toggle options | |
| def toggle_seq_input(choice): | |
| if choice == "protein length": | |
| return gr.update(visible=True, value=None), gr.update(visible=False, value=None) | |
| elif choice == "custom sequence": | |
| return gr.update(visible=False, value=None), gr.update(visible=True, value=None) | |
| def toggle_secondary_structure(choice): | |
| if choice == "sliders": | |
| return gr.update(visible=True, value=None),gr.update(visible=True, value=None),gr.update(visible=True, value=None),gr.update(visible=False, value=None) | |
| elif choice == "explicit": | |
| return gr.update(visible=False, value=None),gr.update(visible=False, value=None),gr.update(visible=False, value=None),gr.update(visible=True, value=None) | |
| # Define the Gradio interface | |
| with gr.Blocks(theme='ParityError/Interstellar') as demo: | |
| gr.Markdown(f"""# Protein Generation via Diffusion in Sequence Space""") | |
| with gr.Row(): | |
| with gr.Column(min_width=500): | |
| gr.Markdown(f""" | |
| ## How does it work?\n | |
| --- [PREPRINT](https://biorxiv.org/content/10.1101/2023.05.08.539766v1) --- | |
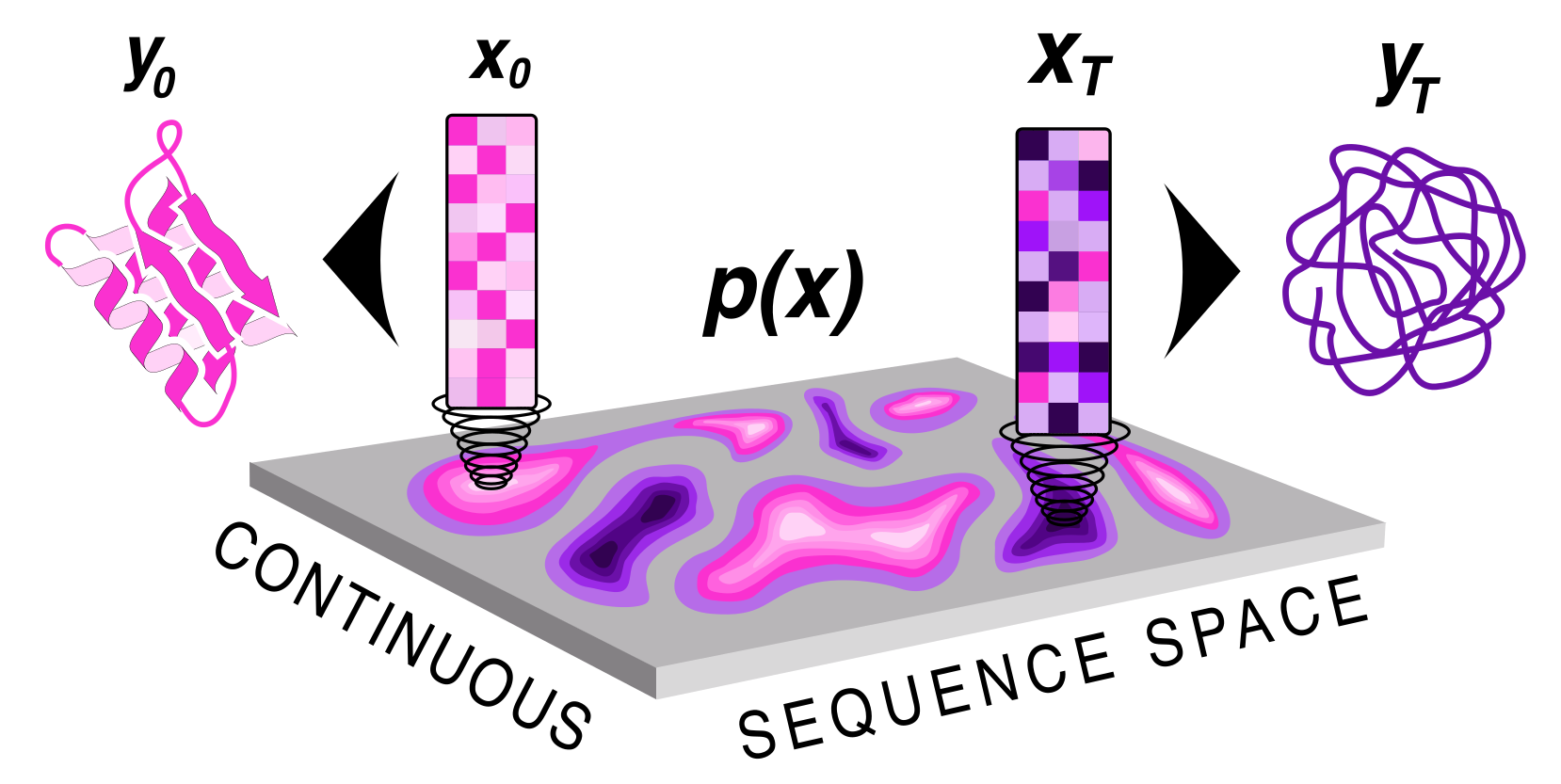
| Protein sequence and structure co-generation is a long outstanding problem in the field of protein design. By implementing [ddpm](https://arxiv.org/abs/2006.11239) style diffusion over protein seqeuence space we generate protein sequence and structure pairs. Starting with [RoseTTAFold](https://www.science.org/doi/10.1126/science.abj8754), a protein structure prediction network, we finetuned it to predict sequence and structure given a partially noised sequence. By applying losses to both the predicted sequence and structure the model is forced to generate meaningful pairs. Diffusing in sequence space makes it easy to implement potentials to guide the diffusive process toward particular amino acid composition, net charge, and more! Furthermore, you can sample proteins from a family of sequences or even train a small sequence to function classifier to guide generation toward desired sequences. | |
|  | |
| ## How to use it?\n | |
| A user can either design a custom input sequence to diffuse from or specify a length below. To scaffold a sequence use the following format where X represent residues to diffuse: XXXXXXXXSCIENCESCIENCEXXXXXXXXXXXXXXXXXXX. You can even design a protein with your name XXXXXXXXXXXXNAMEHEREXXXXXXXXXXXXX! | |
| ### Acknowledgements\n | |
| Thank you to Simon Dürr and the Hugging Face team for setting us up with a community GPU grant! | |
| """) | |
| gr.Markdown(""" | |
| ## Model in Action | |
|  | |
| """) | |
| with gr.Row().style(equal_height=False): | |
| with gr.Column(): | |
| with gr.Tabs(): | |
| with gr.TabItem("Inputs"): | |
| gr.Markdown("""## INPUTS""") | |
| gr.Markdown("""#### Start Sequence | |
| Specify the protein length for complete unconditional generation, or scaffold a motif (or your name) using the custom sequence input""") | |
| seq_opt = gr.Radio(["protein length","custom sequence"], label="How would you like to specify the starting sequence?", value='protein length') | |
| sequence = gr.Textbox(label="custom sequence", lines=1, placeholder='AMINO ACIDS: A,C,D,E,F,G,H,I,K,L,M,N,P,Q,R,S,T,V,W,Y\n MASK TOKEN: X', visible=False) | |
| seq_len = gr.Slider(minimum=5.0, maximum=250.0, label="protein length", value=100, visible=True) | |
| seq_opt.change(fn=toggle_seq_input, | |
| inputs=[seq_opt], | |
| outputs=[seq_len, sequence], | |
| queue=False) | |
| gr.Markdown("""### Optional Parameters""") | |
| with gr.Accordion(label='Secondary Structure',open=True): | |
| gr.Markdown("""Try changing the sliders or inputing explicit secondary structure conditioning for each residue""") | |
| sec_str_opt = gr.Radio(["sliders","explicit"], label="How would you like to specify secondary structure?", value='sliders') | |
| secondary_structure = gr.Textbox(label="secondary structure", lines=1, placeholder='HELIX = H STRAND = S LOOP = L MASK = X(must be the same length as input sequence)', visible=False) | |
| with gr.Column(): | |
| helix_bias = gr.Slider(minimum=0.0, maximum=0.05, label="helix bias", visible=True) | |
| strand_bias = gr.Slider(minimum=0.0, maximum=0.05, label="strand bias", visible=True) | |
| loop_bias = gr.Slider(minimum=0.0, maximum=0.20, label="loop bias", visible=True) | |
| sec_str_opt.change(fn=toggle_secondary_structure, | |
| inputs=[sec_str_opt], | |
| outputs=[helix_bias,strand_bias,loop_bias,secondary_structure], | |
| queue=False) | |
| with gr.Accordion(label='Amino Acid Compositional Bias',open=False): | |
| gr.Markdown("""Bias sequence composition for particular amino acids by specifying the one letter code followed by the fraction to bias. This can be input as a list for example: W0.2,E0.1""") | |
| with gr.Row(): | |
| aa_bias = gr.Textbox(label="aa bias", lines=1, placeholder='specify one letter AA and fraction to bias, for example W0.1 or M0.1,K0.1' ) | |
| aa_bias_potential = gr.Textbox(label="aa bias scale", lines=1, placeholder='AA Bias potential scale (recomended range 1.0-5.0)') | |
| ''' | |
| with gr.Accordion(label='Charge Bias',open=False): | |
| gr.Markdown("""Bias for a specified net charge at a particular pH using the boxes below""") | |
| with gr.Row(): | |
| target_charge = gr.Textbox(label="net charge", lines=1, placeholder='net charge to target') | |
| target_ph = gr.Textbox(label="pH", lines=1, placeholder='pH at which net charge is desired') | |
| charge_potential = gr.Textbox(label="charge potential scale", lines=1, placeholder='charge potential scale (recomended range 1.0-5.0)') | |
| ''' | |
| with gr.Accordion(label='Hydrophobic Bias',open=False): | |
| gr.Markdown("""Bias for or against hydrophobic composition, to get more soluble proteins, bias away with a negative target score (ex. -5)""") | |
| with gr.Row(): | |
| hydrophobic_target_score = gr.Textbox(label="hydrophobic score", lines=1, placeholder='hydrophobic score to target (negative score is good for solublility)') | |
| hydrophobic_potential = gr.Textbox(label="hydrophobic potential scale", lines=1, placeholder='hydrophobic potential scale (recomended range 1.0-2.0)') | |
| with gr.Accordion(label='Diffusion Params',open=False): | |
| gr.Markdown("""Increasing T to more steps can be helpful for harder design challenges, sampling from different distributions can change the sequence and structural composition""") | |
| with gr.Row(): | |
| num_steps = gr.Textbox(label="T", lines=1, placeholder='number of diffusion steps (25 or less will speed things up)') | |
| noise = gr.Dropdown(['normal','gmm2 [-1,1]','gmm3 [-1,0,1]'], label='noise type', value='normal') | |
| with gr.TabItem("Motif Selection"): | |
| gr.Markdown("""### Motif Selection Preview""") | |
| gr.Markdown('Contigs explained: to grab residues (seq and str) on a pdb chain you will provide the chain letter followed by a range of residues as indexed in the pdb file for example (A3-10) is the syntax to select residues 3-10 on chain A (the chain always needs to be specified). To add diffused residues to either side of this motif you can specify a range or discrete value without a chain letter infront. To add 15 residues before the motif and 20-30 residues (randomly sampled) after use the following syntax: 15,A3-10,20-30 commas are used to separate regions selected from the pdb and designed (diffused) resiudes which will be added. ') | |
| pdb_id_code = gr.Textbox(label="PDB ID", lines=1, placeholder='INPUT PDB ID TO FETCH (ex. 1DPX)', visible=True) | |
| contigs = gr.Textbox(label="contigs", lines=1, placeholder='specify contigs to grab particular residues from pdb ()', visible=True) | |
| gr.Markdown('Using the same contig syntax, seq or str of input motif residues can be masked, allowing the model to hold strucutre fixed and design sequence or vice-versa') | |
| with gr.Row(): | |
| seq_mask = gr.Textbox(label='seq mask',lines=1,placeholder='input residues to mask sequence') | |
| str_mask = gr.Textbox(label='str mask',lines=1,placeholder='input residues to mask structure') | |
| preview_viewer = gr.HTML() | |
| rewrite_pdb = gr.File(label='PDB file') | |
| preview_btn = gr.Button("Preview Motif") | |
| with gr.TabItem("MSA to PSSM"): | |
| gr.Markdown("""### MSA to PSSM Generation""") | |
| gr.Markdown('input either an MSA or PSSM to guide the model toward generating samples within your family of interest') | |
| with gr.Row(): | |
| fasta_msa = gr.File(label='MSA') | |
| input_pssm = gr.File(label='PSSM (.csv)') | |
| pssm = gr.File(label='Generated PSSM') | |
| pssm_view = gr.Plot(label='PSSM Viewer') | |
| pssm_gen_btn = gr.Button("Generate PSSM") | |
| btn = gr.Button("GENERATE") | |
| #with gr.Row(): | |
| with gr.Column(): | |
| gr.Markdown("""## OUTPUTS""") | |
| gr.Markdown("""#### Confidence score for generated structure at each timestep""") | |
| plddt_plot = gr.Plot(label='plddt at step t') | |
| gr.Markdown("""#### Output protein sequnece""") | |
| output_seq = gr.Textbox(label="sequence") | |
| gr.Markdown("""#### Download PDB file""") | |
| output_pdb = gr.File(label="PDB file") | |
| gr.Markdown("""#### Structure viewer""") | |
| output_viewer = gr.HTML() | |
| gr.Markdown("""### Don't know where to get started? Click on an example below to try it out!""") | |
| gr.Examples( | |
| [["","125",0.0,0.0,0.2,"","","","20","normal",'','','',None,'','',None], | |
| ["","100",0.0,0.0,0.0,"","W0.2","2","20","normal",'','','',None,'','',None], | |
| ["","100",0.0,0.0,0.0, | |
| "XXHHHHHHHHHXXXXXXXHHHHHHHHHXXXXXXXHHHHHHHHXXXXSSSSSSSSSSSXXXXXXXXSSSSSSSSSSSSXXXXXXXSSSSSSSSSXXXXXXX", | |
| "","","25","normal",'','','',None,'','',None], | |
| ["XXXXXXXXXXXXXXXXXXXXXXXXXIPDXXXXXXXXXXXXXXXXXXXXXXPEPSEQXXXXXXXXXXXXXXXXXXXXXXXXXXIPDXXXXXXXXXXXXXXXXXXX", | |
| "",0.0,0.0,0.0,"","","","25","normal",'','','',None,'','',None], | |
| ["","",0.0,0.0,0.0,"","","","25","normal",'','', | |
| '9,D10-11,8,D20-20,4,D25-35,65,D101-101,2,D104-105,8,D114-116,15,D132-138,6,D145-145,2,D148-148,12,D161-161,3', | |
| './tmp/PSSM_lysozyme.csv', | |
| 'D25-25,D27-31,D33-35,D132-137', | |
| 'D26-26','./tmp/150l.pdb'] | |
| ], | |
| inputs=[sequence, | |
| seq_len, | |
| helix_bias, | |
| strand_bias, | |
| loop_bias, | |
| secondary_structure, | |
| aa_bias, | |
| aa_bias_potential, | |
| #target_charge, | |
| #target_ph, | |
| #charge_potential, | |
| num_steps, | |
| noise, | |
| hydrophobic_target_score, | |
| hydrophobic_potential, | |
| contigs, | |
| pssm, | |
| seq_mask, | |
| str_mask, | |
| rewrite_pdb], | |
| outputs=[output_seq, | |
| output_pdb, | |
| output_viewer, | |
| plddt_plot], | |
| fn=protein_diffusion_model, | |
| ) | |
| preview_btn.click(get_motif_preview,[pdb_id_code, contigs],[preview_viewer, rewrite_pdb]) | |
| pssm_gen_btn.click(get_pssm,[fasta_msa,input_pssm],[pssm_view, pssm]) | |
| btn.click(protein_diffusion_model, | |
| [sequence, | |
| seq_len, | |
| helix_bias, | |
| strand_bias, | |
| loop_bias, | |
| secondary_structure, | |
| aa_bias, | |
| aa_bias_potential, | |
| #target_charge, | |
| #target_ph, | |
| #charge_potential, | |
| num_steps, | |
| noise, | |
| hydrophobic_target_score, | |
| hydrophobic_potential, | |
| contigs, | |
| pssm, | |
| seq_mask, | |
| str_mask, | |
| rewrite_pdb], | |
| [output_seq, | |
| output_pdb, | |
| output_viewer, | |
| plddt_plot]) | |
| demo.queue() | |
| demo.launch(debug=True) | |